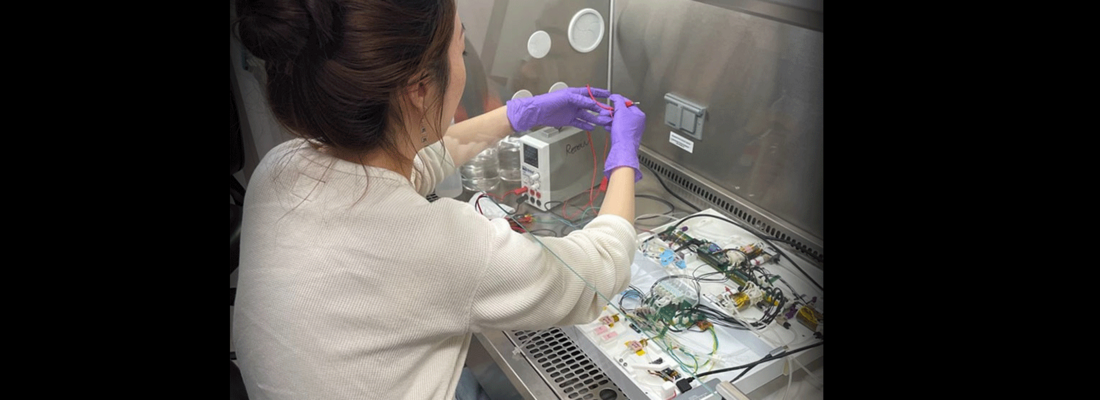
The experiment involves collaboration with MIT Media Lab Space Exploration Initiative, Harvard Medical School, and Seed Health, and is funded in part by the National Renewable Energy Laboratory (NREL)-led Bio-Optimized Technologies to keep Thermoplastics out of Landfills and the Environment (BOTTLE™) Consortium, which is sponsored by the Department of Energy’s Bioenergy Technologies Office and Advanced Materials and Manufacturing Technologies Office. BOTTLE researchers from NREL working on this project include synthetic biologist Allison Werner, biochemists Erika Erickson and Natasha Murphy, analytical chemists Kelsey Ramirez and Morgan Ingraham, and BOTTLE CEO Gregg Beckham.
The two samples—PETase and P. putida—were stowed within a custom bio-experimentation payload inside the SpX-26 resupply mission, which departed from Kennedy Space Center on Nov. 26. Together, the enzyme and microbe provide a solution for upcycling polyethylene terephthalate (PET), a popular polyester used in clothes, bottles, and more.
“This will be a demonstration of upcycling plastics to new materials in low Earth orbit,” said Allison Werner, a synthetic biologist and the project lead at NREL. “Broadly, the experiment aims to accomplish three things: first, to flight-test a new autonomous cultivation system that expands the capabilities for unmanned experiments; second, to assess the effect of spaceflight on enzymatic depolymerization of PET plastic; and third, to investigate the genome and proteome of bacteria engineered to convert depolymerized PET to a nylon precursor.”
This upcycling pathway begins with the PETase enzyme breaking down PET plastic into its precursor, terephthalic acid, a chemical that is produced from petroleum at several million tons per year. Terephthalic acid is then fed to P. putida, a common soil bacteria with pathways engineered to consume this compound. The bacteria convert the acid into an enhanced precursor for nylon, thus converting a waste plastic into building blocks for a high-performance plastic. But to carry out this cycle on the ISS, the project team needed to invent an autonomous experiment-in-a-box, and their solution could allow a new approach to microbial studies in space and other remote sites.
Autonomous Culturing for Remote Experiments
The project team wanted to find a solution for performing biological research without hands-on help in space. Ben Fram, a Ph.D. student at Harvard Medical School, is one of the project researchers contributing to this autonomous experiment.
“Biological experiments require a lot of hand-holding. Cells need to be passaged into new media, and enzyme reactions need refreshing,” Fram said. “Without an astronaut’s help, experimental options on the ISS are limited.”
To address this need, the team invented a self-contained experiment with timed sampling, passaging, and automatic data logging. The autonomous culturing system is developed by researchers from MIT Media Lab Space Exploration Initiative, including Xin Liu, a research lead on the payload system. Liu describes it as a compact, modular, spaceflight-ready bio-culturing system with its accompanying protocol that enables continuous microbial cultivation and adaptable protocol execution without human intervention.
“It is crucial for us to work with biologists from the beginning so that we can design the system with biology at its center and make it flexible enough to accommodate different requirements as the research goals evolve,” Liu said. “We hope this system can be foundational work for more accessible biological research for human spaceflights.”
The bio-culturing payload design is intended to be open source and will be fully explained in a forthcoming publication. It uses hardware that can be purchased from commercial vendors and parts that can be 3D-printed by anyone, and it is built to accommodate biological research in a closed system. With minor modifications, it could potentially be used in other remote research, like culturing in the ocean or polar ice.
First, bacteria are reconstituted from powder, and PETase is transferred into new buffer. They are then moved between chambers to be continually cultivated and sampled. Pumps inject the chambers with media—plastics for the enzymes and acid for the bacteria—allowing reactions to take place and cells to grow. At timed intervals, the box saves samples by autonomously pumping the solutions into preservation bags, so that growth and chemical products are benchmarked along the way. The box will also download analysis data to be reviewed in the labs. And on Earth, the exact same experiment is underway for comparison.
Learning From Space Mutants
Without knowing exactly what will change for bacteria in space, the researchers have reason to expect the terrestrial and microgravity results to be different. Past experiments have shown that weightlessness affects metabolism, gene expression, and protein regulation in microorganisms, but the effect on the team’s PET-upcycling bacteria is unknown. The effect of microgravity on interfacial biocatalysis—enzymes degrading sheets of PET—is similarly unknown.
“We know that spaceflight will have effects from radiation and microgravity, but we do not have a priori knowledge of the effect of these stressors on the enzyme or engineered strain’s performance,” Werner said.